TECHNOLOGY SEED
- Technology Platforms
Development of Programmable Splice Regulatory RNA To Induce Skipping of Target Exons – Alternatives to Splicing Regulatory Antisense Oligonucleotides
Artificial RNA molecules to regulate mRNA splicing for treating genetic diseases and advancing gene therapy

Source: scienceDISPLAY, stock.adobe.com/uk/427218185, stock.adobe.com
Background
Aberrant mRNA splicing is implicated in the pathogenesis of numerous genetic and neurodegenerative diseases, making precise control of splicing an important therapeutic target. Antisense oligonucleotides (ASOs) have been utilized for splicing modulation; however, they present intrinsic challenges such as chemical modification–related toxicity, limited cellular uptake, and short half-life requiring frequent dosing. Although CRISPR/Cas9 genome editing offers powerful genetic manipulation, its irreversibility, risks of over-editing, and vector size limitations render it less suitable for reversible splicing control. These limitations highlight the need for new RNA-based technologies that enable safe, flexible, and high-precision regulation of gene expression at the RNA level.
Technology Overview
Through a collaborative effort between Hokkaido University and Setsunan University, an artificial RNA molecule has been developed by leveraging the splicing-suppressive mechanism of the non-coding RNA 4.5SH.
This artificial RNA is characterized by two key components:
- A stem–loop structure derived from 4.5SH RNA, which recruits splicing-suppressor effector proteins to the target site; and
- An antisense sequence complementary to the target pre-mRNA, providing target specificity.
By simultaneously recognizing the target exon and recruiting splicing regulatory factors, this artificial RNA efficiently induces exon skipping. The molecule can be used directly as a synthetic nucleic acid, and its structure is also compatible with incorporation into expression vectors or viral vectors such as AAV, allowing flexible consideration of delivery strategies depending on the intended application.
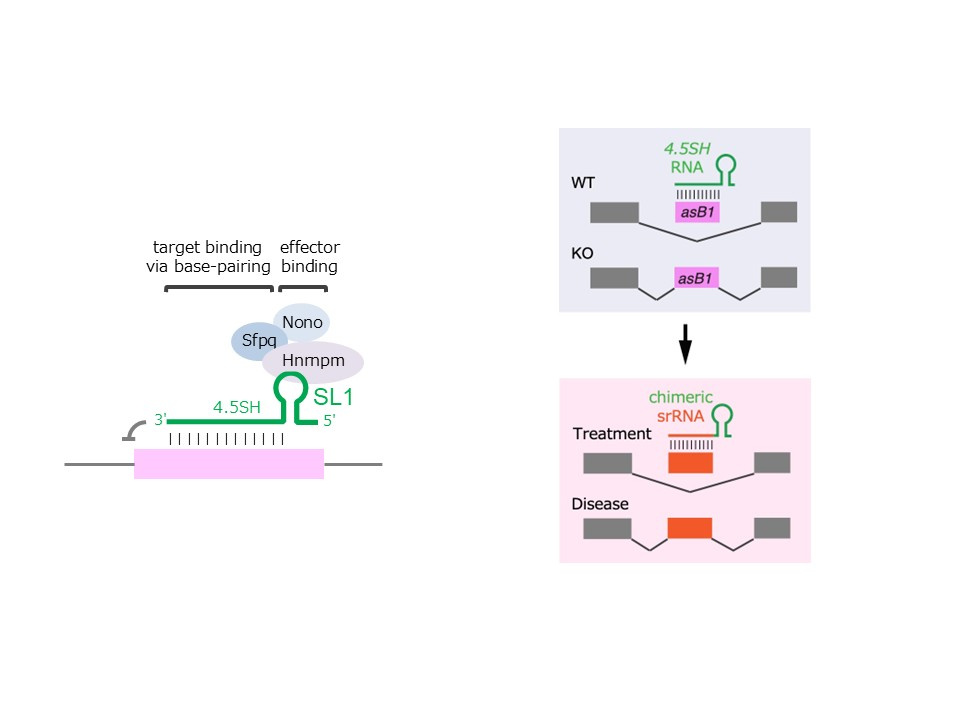
Figure 1: Design concept of programmable splicing regulatory RNA (srRNA) based on 4.5SH structure
This figure shows the design principle of artificial splicing regulatory RNA (srRNA), derived from the 4.5SH RNA architecture. The srRNA consists of two functional modules: an effector-binding module (derived from stem–loop 1) and a target-recognition module complementary to the target exon. By modifying the recognition sequence, srRNA can be programmed to induce exon skipping for specific genes.
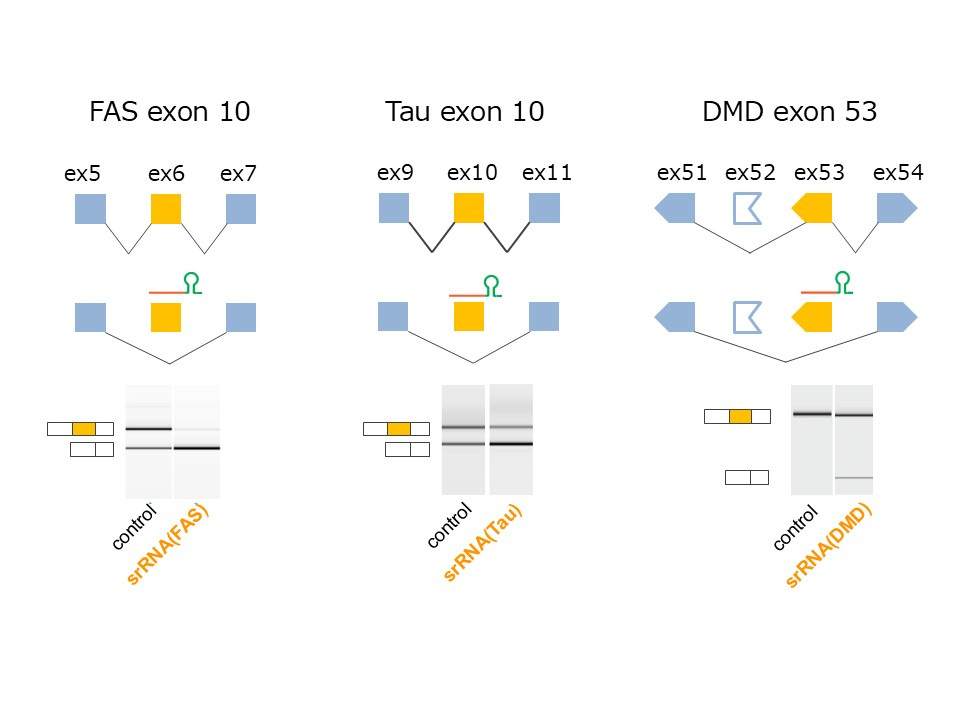
Figure 2: Reporter assay: Programmed srRNAs induce skipping of target exons
Alternative splicing of FAS-ex6, Tau-ex10, and Dmd-ex53 in HEK293 cells transfected with splicing minigenes and expression vectors for the chimeric RNAs. The chimeric RNAs efficiently induced exon skipping.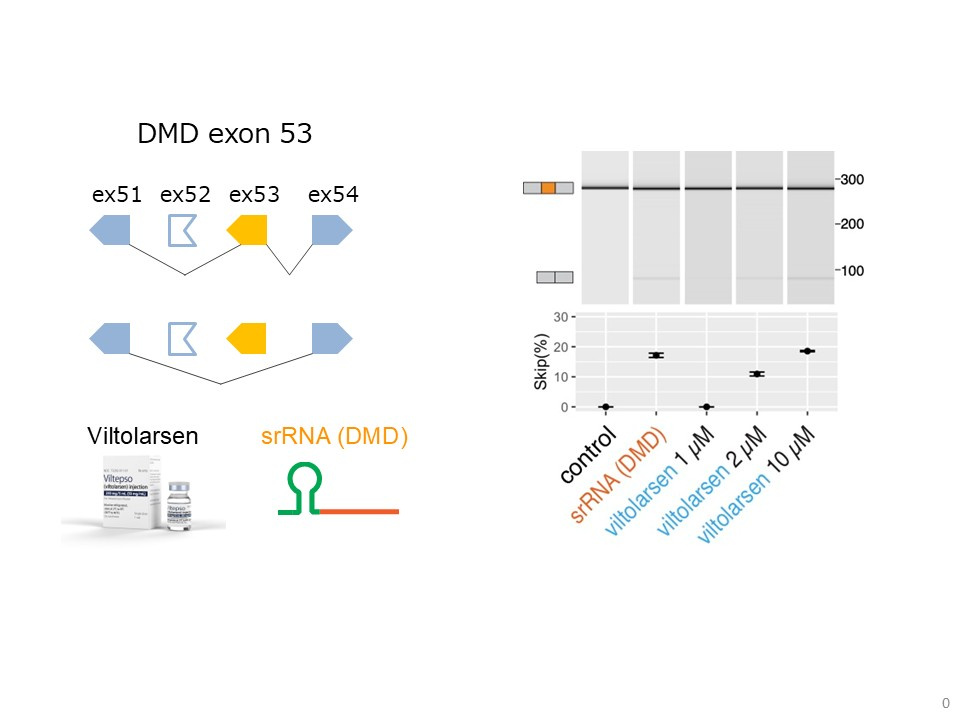
Figure 3: Reporter assay: Plasmid-expressed srRNA (DMD) shows exon-skipping activity comparable to morpholino ASO at the corresponding concentration
Quantification of the alternative splicing of DMD-ex53 in HeLa cells transfected with the splicing minigene, and either vectors expressing the chimeric RNA or different concentrations of Viltolarsen, a morpholino antisense oligonucleotide (ASO) approved for the treatment of Duchenne muscular dystrophy (DMD).
The exon-skipping efficiency of the chimeric RNA was comparable to that observed with 10 µM Viltolarsen, a concentration used in therapeutic applications.
These results indicate that srRNA achieves splicing modulation equivalent to clinically used ASOs, highlighting its potential as a gene-modulatory RNA platform.
Further Details:
Stage of Development
- In vitro validation
- Preclinical stage
Benefits
- A novel mechanism for splicing regulation
- Unlike ASOs, which rely primarily on passive hybridization-based modulation, this technology employs an artificial RNA capable of actively recruiting splicing-suppressor proteins, enabling efficient and selective induction of exon skipping.
- Enhanced structural stability derived from the stem–loop architecture
- Retention of the 4.5SH-based secondary structure contributes to RNA conformational stability and supports sustained functional activity.
- Customizable for multiple target genes
- The platform has demonstrated applicability to several genes—including FAS, DMD, MAPT, and FKTN—supporting its utility for diverse disease-associated splicing events.
- Compatibility with multiple delivery modalities
- The artificial RNA can function as a synthetic nucleic acid and is structurally amenable to incorporation into AAV vectors, plasmid expression systems, or LNP-based delivery technologies, providing flexibility for future formulation development.
Applications
This technology has potential applications across diseases in which aberrant splicing contributes to pathology, particularly those treatable through exon skipping–based approaches.
- Neuromuscular and neurodegenerative diseases:
- Duchenne muscular dystrophy, myotonic dystrophy, ALS, FTDP-17, SMA, familial dysautonomia, Fukuyama congenital muscular dystrophy, hereditary spastic paraplegia, etc.
- Genetic, metabolic, and immune-related disorders
- Thalassemia, cystic fibrosis, Wilson disease, Pompe disease, Rett syndrome, NF2, rheumatoid arthritis, inflammatory disorders, metabolic dysfunction, and others.
- Hematological disorders and malignancies
- MDS, AML, CMML, and other diseases involving splicing-factor dysfunction.
- Neurodevelopmental and sensory disorders
- Huntington’s disease, Alzheimer’s disease, epileptic encephalopathies, Dravet syndrome, retinal degeneration, etc.
- Basic research
- Analysis of alternative splicing mechanisms, construction of RNA-based disease models, and use as a research tool for studying splicing regulators.
Opportunity
The research team are seeking collaborations with biotechnology and pharmaceutical companies for preclinical validation and translational research.
Particularly interested in partnerships with firms engaged in nucleic acid therapeutics development—including antisense oligonucleotide (ASO) programs—as well as those with expertise in vector engineering and nucleic acid delivery technologies.
Licensing and co-development agreements are also welcomed.
Further Opportunity Details:
- Joint development for in vivo delivery optimization and disease model validation
- Collaborative exploration of new indications in RNA therapeutics
- Licensing opportunities for therapeutic applications and platform expansion
Link
inpart
Contact